A. Ice Skating
Notice that the existence of a snow drift at the point (x, y) implies that "if I'm on the horizontal line at y then I am certainly able to get to the vertical line at x, and vice versa". Thus, the snow drifts are the edges of a bipartite graph between x- and y- coordinates. The number of snow drifts that need to be added to make this (as well as the original) graph connected is the number of its connected components reduced by one.
B. Blackboard Fibonacci
If you look at the described process backwards, it resembles the Euclidean algorithm a lot. Indeed, if you rewinded a recording of Bajtek's actions, he always takes the larger out of two numbers (say
Unable to parse markup [type=CF_TEX]
) and replaces them by a - b, b. Since we know one of the final numbers (r) we can simply check all numbers between 1 and r and run a faster version of Euclid's algorithm (one that replaces a, b by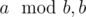 ) for all possibilities for a total runtime of
) for all possibilities for a total runtime of 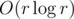 . This was one of the expected solutions.
. This was one of the expected solutions.However, with some insight, it can be seen that this optimization is in fact not neccessary — we can simply simulate the reverse process as described (replacing a, b by a - b, b) for all candidates between 1 and r and the total runtime of our algorithm will remain 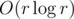 . The proof of this fact is left to the reader.
. The proof of this fact is left to the reader.
C. Formurosa
One of the major difficulties in this problem is finding an easily formulated condition for when Formurosa can be used to distinguish the bacteria. Let Formurosa's digestive process be a function F(s) that maps binary sequences of length m to elements of {0, 1}. It turns out that the condition we seek for can be stated as follows:
We can distinguish all the bacteria if and only if there exists a sequence s of length m for which F(s) ≠ F( - s), where - s is the negation of s.
First, not that if no such sequence exists, then there is no way to distinguish between zero and one. If such a sequence exists, we can pick any two bacteria a and b and try both ways to substitute them for 0 and 1 in the expression. If the two expressions evaluate to different values, we will determine the exact types of both bacteria. Otherwise, we will be certain that the bacteria are of the same type. Repeating the process for all pairs of bacteria will let us identify all the types (since it is guaranteed that not all bacteria are of the same type).
To determine whether such a sequence s exists, dynamic programming over the expression tree of Formurosa can be applied. The model solution keeps track for each subtree G of the expression which of the following sequences can be found:
- a sequence s such that G(s) = G( - s) = 0
- a sequence s such that G(s) = G( - s) = 1
- a sequence s such that G(s) ≠ G( - s)
D. Bitonix' Patrol
Observation 1.
Fuel tanks for which capacity gives the same remainder 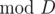 are equivalent for Bitonix's purposes. Moreover, fuel tanks for which the capacities' remainders
are equivalent for Bitonix's purposes. Moreover, fuel tanks for which the capacities' remainders 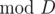 sum to D are also equivalent. Out of every group of equivalent tanks, the agency can only leave at most one.
sum to D are also equivalent. Out of every group of equivalent tanks, the agency can only leave at most one.
Observation 2.
If more than six tanks remain, Bitonix can certainly complete his patrol. Indeed, let us assume that 7 tanks were left undestroyed by the agency. Out of the 128 possible subsets of those tanks, at least two distinct ones, say A and B, sum up to the same remainders modulo D. Thus, if Bitonix moves forward with tanks from A - B and backwards with tanks from B - A, he will finish at some station after an actual journey.
Because of observations 1 and 2, it turns out that a simple recursive search suffices to solve the problem. However, because of the large constraints, it may prove necessary to use some optimizations, such as using bitmasks for keeping track of what distances Bitonix can cover.
E. Alien DNA
Note that it is easy to determine, looking at only the last mutation, how many letters it adds to the final result. Indeed, if we need to print out the first k letters of the sequence, and the last mutation is [l, r], it suffices to find out the length of the overlap of segments [1, k] and [r + 1, 2r - l + 1]. Say that it is x. Then, after the next to last mutation, we are only interested in the first k - x letters of the result — the rest is irrelevant, as it will become "pushed out" by the elements added in the last mutation. Repeating this reasoning going backwards, we shall find out that we can spend linear time adding letters to the result after every mutation, which turns out to be the main idea needed to solve the problem.
For a neat O(n2 + k) implementation of this idea you can check out ACRush's solution: 2029369.
The only other contestant to solve the problem during the competition, panyuchao, used a slightly different approach, based in part on the same idea. Check out his ingenious, astonishingly short 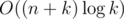 solution here: 2028007.
solution here: 2028007.










better late than never
nice
What is wrong in my solution of problem A... 11962492
What's the speciality with test case #12? I seem to get WA here.
Could you please share your solution? I'm getting WA on test case #12
Here's my solution: My solution
Nevermind, problem solved :B
same problem with me !!! how did you escaped from it ?
Nevermind, problem solved. :]
Very Nice Question A
For QUES A.
For those who couldn't pass test case #12. Try this and you may realize the error.
3
1 2
3 4
3 2
The solution to Airport question (https://codeforces.com/contest/218/submission/211784519) 211784519
can you provide explaination